(C) 2014 Gerhard Rambold. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 4.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Over the past 12 years, the lichen trait database LIAS light as a component of the LIAS information system, has grown to a considerable pool of descriptive data based on 71 different qualitative, quantitative, and text characters, for nearly 10.000 lichen taxa, being phylogenetically arranged according to the MycoNet classification. It includes information on morphological, ecological and chemical traits. Multilinguality or internationalization options have become a central challenge of the project. At present, 18 language versions of the database and web interface exist. LIAS light data are accessible in DELTA format and to be used locally and web browser-based, via NaviKey applet.
Identification, trait database, characters, biodiversity informatics
For the purpose of giving access to treated taxa, monographic and floristic treatments are often assigned with traditional dicho- or polytomous identification keys. Until today, such keys have proven to be of highest value as they mostly provide identification criteria optimized for a specific set of taxa or an area. Nevertheless, the growing use of the internet as well as the need of open access to information and knowledge on biodiversity at a higher or global scale has led to an increased use of internet-related technologies for identifying taxa (
Over the past 12 years, the lichen trait database LIAS light (http://liaslight.lias.net ) has grown to a considerable pool of descriptive data for lichens with information on morphological, ecological and chemical traits. It is part of the LIAS data platform, which has been initiated as a web project in 1995 (
There are two other parts of the site (LIAS main and LIAS checklists) which have been frozen in favour of LIAS light which works based on a reduced number of descriptors (see below) and will incorporate in the near future occurrence data at the country level as well. While the concept of LIAS main was based on lichenologist community driven data input, entry of LIAS light data was primarily taken over by laboratory members of the first author, starting with compilation data from outstanding lichen floras like
The number of taxa is now close to 10, 000. Estimate numbers of accepted names of lichens at a global scale range between 13, 500 and 20, 000. Given the number of 17, 000 described, accepted lichen species of the world as proposed by
The taxon names used for the LIAS light dataset are maintained in LIAS names. For most of the records, modern taxonomic concepts are applied when literature references exist. The higher classification backbone is based on MycoNet data (http://fieldmuseum.org/explore/myconet ) and updated accordingly. By activating the resulting LIAS light taxon names, access to additional information and links to further resources is provided. Aside form LIAS names (see above), they include a) an interface using the web service of Recent Literature of Lichens (http://www.bio.uni-bayreuth.de/liaslight/service/rll_wrap.php ); b) a lichen image accession interface (http://www.bio.uni-bayreuth.de/liaslight/service/LichenImages.php ) to reliable web sites with significant images; and c) several less specific portals like Wikispecies, Google Images, NCBI Search and others.
Associated with LIAS light is LIAS metabolites (http://liaslight.lias.net/Identification/Navikey/Metabolites/index.html ), a database compiled by J. A. Elix with additions of K. Kalb (Neumarkt, Germany). This LIAS database component includes the values of TLC, HPLC and MS for about 880 secondary lichen compounds, together with additional information like the substance class and names of related metabolites. It is useful for interpreting chromatographic and spectroscopic analysis results. Soon each data record of LIAS metabolites will be supplemented with the chemical formulas and further representations including SMILE codes.
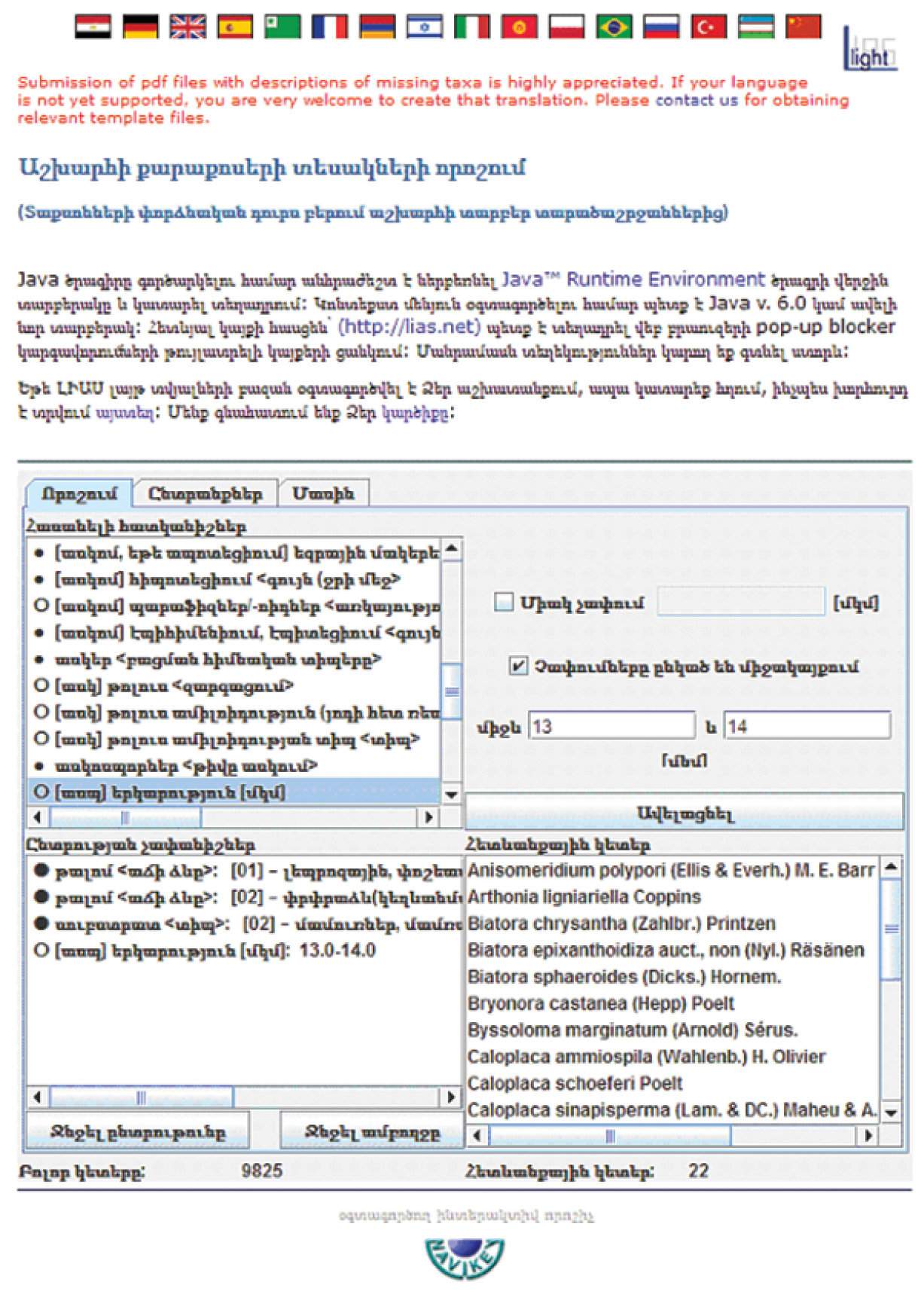
Multilinguality or internationalization options have become a central challenge of the LIAS light project. At present, versions in 18 languages exist: aside from English, being Arabic, Armenian (Fig. 1), Chinese, Esperanto, Farsi, French, German, Hebrew, Italian, Korean, Kyrgyz, Polish, Portuguese, Romanian, Russian, Uzbek, Spanish, and Turkish. The translation of the subject-specific vocabulary is intended as a contribution of preventing ‘lingual erosion’ in the languages aside from English. Regarding the 18 languages actually included in LIAS light, it becomes apparent that several ones (like French, German, Polish, Russian, Spanish etc.) easily cover all general and specific terminologies, while others (like Kyrgyz or Chinese) are only capable of providing the general terms but not or much less the specific ones, (e.g. the scientific names of secondary metabolites). In such cases, supplementation of the respective character state list by terms from English or Russian (in the case of Kyrgyz) or another widely spoken language is required. The use of a native language is desired, especially by students as well as amateur communities, who are involved in monitoring projects in many countries of the world. The translation of the LIAS light character list and the NaviKey web interface contents into the various languages have been achieved by colleagues in biology or lichenology (http://liaslight.lias.net/About/Impressum.html ).
LIAS light web interface NaviKey in Armenian language.
LIAS light content is provided via NaviKey, a Java applet and application for accessing descriptive data (http://www.navikey.net/ ). It provides quite a range of possibilities to modify the queries for obtaining improved results. One of the most important option is ‘Retain items unrecorded for the selected characters’ being a switch between a ‘data query mode’ (checkbox “set off”, as the default) and an ‘identification mode’ (checkbox “set on”). While the query mode is strict and only provides the directly matching data according to the selection, the identification mode is more tolerant in the way that it matches positively also in the case, that a respective character state is not specified in the respective data record. During identification, the option to simultaneously select more than one character state, either with OR (‘Retain matching at least one selected state’ checkbox “set on”, as the default) or with AND (checkbox “set off”) selection criteria, facilitates one making decisions when specifying states of categorical characters. For instance, the decision to select certain colour nuances may be relatively difficult. However, switching between the two modes is not yet possible during one and the same identification procedure, but will be implemented in the future.
The LIAS light database with its NaviKey web interface still exhibits certain shortcomings and does not fully provide satisfactory results in all cases. Nevertheless, it is foreseeable that by continued data entry and revision and with some improvements of the user interface, LIAS light will become a feasible tool for lichen identification. Especially the intended inclusion of country checklist information from reliable original sources (e.g., the original checklists of Austria, Australia, France, Germany, or North America), will entail improved resolution of data and therefore more reliable identification results, especially for countries with a more or less saturated knowledge of taxon occurrence. The use of checklist information as a prefilter function also beneath country level is being considered. Since October 2013, GBIF offers the option of user-defined filter creation for dynamically generating spatially defined species lists. By implementing access to this service LIAS light will be enabled to use this additional option of taxon preselection. For the next future, functionality of the NaviKey software will be extended by enabling the dynamic generation of taxon description web pages. LIAS metabolites will be optimized and new languages will be added on request.
All contents of the lingual versions are downloadable under a Creative Commons Attribution-Non-Commercial-No Derivative Works 3.0 Unported License as archive data files from the respective LIAS light language version page (e.g., http://liaslight.lias.net/Identification/Navikey/data/download/LIASlightDataFilesWorld.pl_PL.zip for the data set in the Polish language). The data may be evoked by a local installation of NaviKey and be even used offline, which, for instance, may be of value for work in remote areas with low bandwidth internet. Server logs indicate an increasing acceptance of LIAS light. The relative number of re-visitors or users, respectively, is steadily growing and is a clear argument in favour of internationalizing the project – there is a striking correlation between visitor location at the country level and the language version being selected. A citation of LIAS light by users in their publications is highly appreciated and should be in accordance with the imprint page (http://liaslight.lias.net/About/Impressum.html ).
For translating the LIAS light database schema and user interface into the various languages, we would like to thank Martin Brüggemeier, Ahaus, Germany (Esperanto), Tiziana Cossu, Sassari, Italy (Italian), Evgeny A. Davydov, Barnaul, Russia (Russian), Decebal-Radu Ciurchea, Cluj-Napoca, Romania (Romanian), Estelle Enaud, Louvain-la-Neuve, Belgium (French), Arsen Gasparyan, Yerevan, Armenia (Armenian), Iris Gonzáles, Managua, Nicaragua (Portuguese), Yongdoo Kim, Gwangju, South Korea (Korean), Tamar Kis-Papo, Haifa, Israel (Hebrew), Nathalie Moske, Bayreuth, Germany (Turkish), Gulnara A. Muhambetalieva, Karakol, Kyrgyzstan, (Kyrgyz, Uzbek), Nicole Nöske, Berlin, Germany (German), Piotr Jan Osyczka, Krakow, Poland (Polish), Claude Roux, Mirabeau, France (Esperanto, French), Veronica Sanz, Barcelona, Spain (Spanish), Ayhan Şenkardeşler, Izmir, Turkey (Turkish), Sameh Mohammad Shaddad, Sassari, Italy (Arabic), Mohammad Sohrabi, Helsinki, Finland and Tehran, Iran (Armenian, Farsi, Turkish), Ileana Téllez, Managua, Nicaragua (Portuguese), Reinaldo Vargas Castillo, Concepción, Chile (Spanish), Jiang-Chun Wei, Beijing, China (Chinese), and Xin Li Wei, Beijing, China (Chinese).
The work was supported in part by the Federal Ministry of Education and Research, Germany (BMBF) under project ID 01 LI 1001 B ‘GBIF-D’ and by the German Research Foundation (DFG) as a project of the LIS infrastructure program grants (project IDs INST 747/1-1, RA 731/11-2, and TR 290/5-1) and in the context of the funding initiative DFG/CNPq: German-Brasilian consultative visits under DFG project ID 444 BRA – 113 / 58 / 0-1. Support was also granted by the European Union’s 7 Framework Programme (project ID FP7/2007-2013) in the 4D4Life project context (grant agreement ID 238988).