(C) 2013 Steven D. Leavitt. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
Recent studies using sequence data from multiple loci and coalescent-based species delimitation have revealed several species-level lineages within the phenotypically circumscribed taxon Rhizoplaca melanophthalma sensu lato. Here, we formally describe five new species within this group, Rhizoplaca occulta, Rhizoplaca parilis, Rhizoplaca polymorpha, Rhizoplaca porterii, and Rhizoplaca shushanii, using support from the coalescent-based species delimitation method implemented in the program Bayesian Phylogenetics and Phylogeography (BPP) as the diagnostic feature distinguishing new species. We provide a reference DNA sequence database using the ITS marker as a DNA barcode for identifying species within this complex. We also assessed intraspecific genetic distances within the six Rhizoplaca melanophthalma sensu lato species. While intraspecific genetic distances within the five new species were less than or equal to the lowest interspecific pairwise comparison values, an overlap in genetic distances within the Rhizoplaca melanophthalma sensu stricto clade suggests the potential for additional phenotypically cryptic lineages within this broadly distributed lineage. Overall, our results demonstrate the potential for accurately identifying species within the Rhizoplaca melanophthalma group by using molecular-based identification methods.
Bio-monitoring, BPP, coalescence, cosmopolitan distribution, cryptic species, molecular identification, symbiosis
The lichen symbiosis has been highly successful with more than 18, 000 currently accepted species of lichen-forming fungi (
Using DNA barcoding, a reference DNA sequence database generated from expertly identified specimens can provide an effective alternative to phenotype-based identification of lichen-forming fungal species (
Rhizoplaca Zopf, as currently circumscribed, is a morphologically diverse, polyphyletic genus (
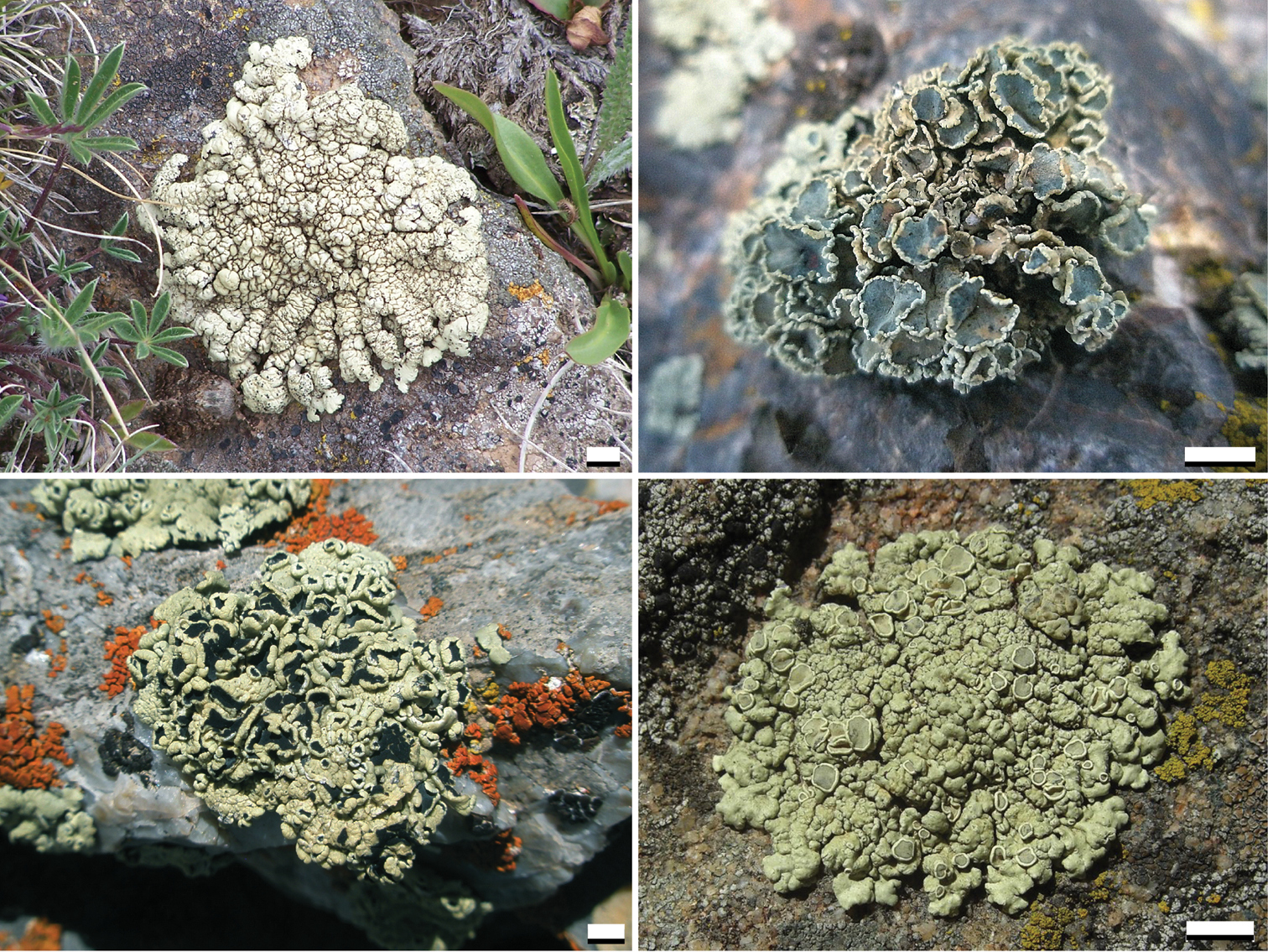
Within Rhizoplaca melanophthalma s. l. (Fig. 1),
Variation in morphology and habit within Rhizoplaca melanophthalma sensu lato. Scale bar = 5 mm.
While morphological and chemical character differences have traditionally served as proxies for identifying reproductively isolated groups, multilocus coalescent-based species delimitation methods can provide a more direct assessment of gene flow and independent lineage status through genetic analysis. Coalescent-based methods can provide a more direct and replicable approach for assessing hypotheses of evolutionary independence, regardless of whether putative lineages differ in potentially subjective phenotypic character systems (
In this paper we use support from the coalescent-based species delimitation method implemented in the program BPP (
Rhizoplaca melanophthalma s.l. is frequently used in air quality bio-monitoring studies (
Using multilocus sequence data generated from Rhizoplaca specimens collected throughout the Intermountain region of western North America,
In this study, we used the ITS alignment and phylogeny reported in
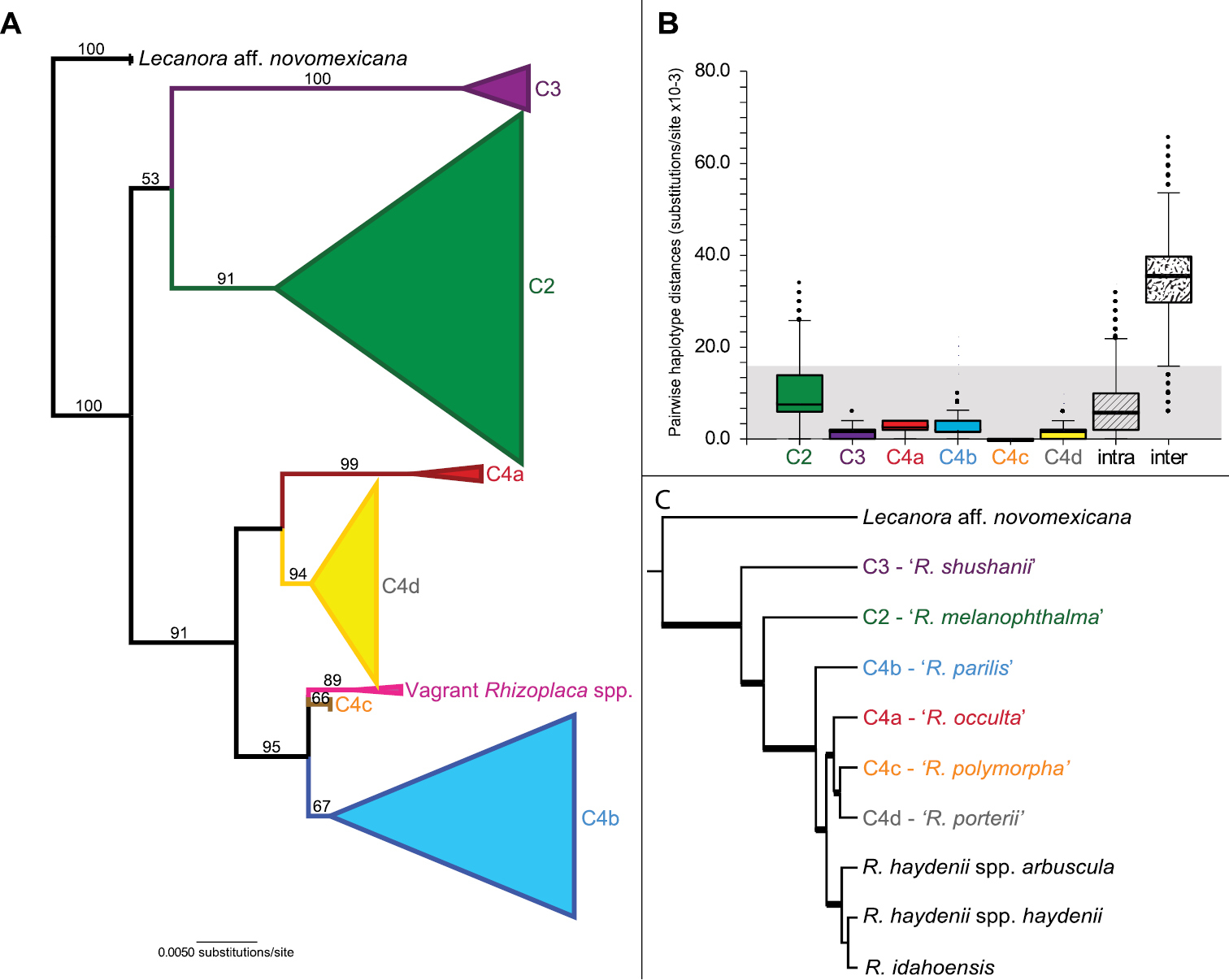
A Cartoon representation of the maximum likelihood ITS topology obtained from 240 Rhizoplaca melanophthalma sensu lato specimens in Leavitt et al. (in review). Values at each node indicate non-parametric-bootstrap support; only support values > 50% are indicated B Box plots of ITS genetic distances within each new species, all intraspecific distances, and all interspecific distances. In each box plot, the box shows the interquartile range (IQR) of the data. The IQR is defined as the difference between the 75th percentile and the 25th percentile. The solid and dotted line through the box represent the median and the average length, respectively; and C The coalescent-based species-tree for the Rhizoplaca melanophthalma species-complex estimated from five genetic markers (ITS, IGS, group I intron, β-tubulin, and MCM7 loci) in
In the present study, we calculated pairwise distances to characterize both intra- and interspecific variation within and among candidate species-level lineages. Pairwise distances can be viewed as a rough measure for the overall sequence divergence (
The Barcode of Life Data Systems (BOLD;
We assessed the utility of the ITS region for BLAST-based specimen identification in the Rhizoplaca melanophthalma group by conducting searches of the newly reported ITS sequences in
The marginal posterior probability of speciation (speciation probability) was estimated in
For each candidate species, the pairwise distances among the different ITS haplotypes were estimated and the distribution of distances plotted (Fig. 2B). Intraspecific genetic distances in candidate species-level lineages ‘C3’, ‘C4a’, ‘C4b’, ‘C4c’ and ‘C4d’ were less than or equal to the lowest interspecific pairwise comparison values (0.006 substitutions/site), with the exception of outlier values in ‘C4b’ and ‘C4d’ (Fig. 2B). The outlier values within clade ‘C4b’ were based on comparisons from two sequences retrieved from GenBank (EF095278 and EF095287). We were unable to verify the quality of these sequences. Within candidate lineage ‘C2’, intraspecific values largely fell below the lower quartile of interspecific pairwise comparison values (Fig. 2B), although there was substantial overlap between intra- and interspecific genetic distance values. Mean distance values, standard deviations and the range of intraspecific distances within the six candidate species are reported in Table 1.
Mean genetic distance values (given as number of nucleotide substitution per site) and range of intraspecific distances for the Rhizoplaca melanophthalma species-complex. Numbers within parentheses indicate the number of sampled individuals/lineage; and values following mean genetic distance represent standard deviations.
| Species/Clade | Mean | range |
|---|---|---|
| Rhizoplaca melanophthalma ‘C2’ (98) | 0.009 ± 0.006 | 0.0 – 0.034 |
| Rhizoplaca occulta ‘C4a’ (5) | 0.003 ± 0.001 | 0.0 – 0.004 |
| Rhizoplaca parilis ‘C4b’ (56) | 0.003 ± 0.004 | 0.0 – 0.022 |
| Rhizoplaca polymorpha ‘C4c’ (6) | 0.0 ± 0 | 0.0 – 0.0 |
| Rhizoplaca porterii ‘C4d’ (57) | 0.002 ± 0.002 | 0.0 – 0.010 |
| Rhizoplaca shushanii ‘C3’ (13) | 0.001 ± 0.002 | 0.0 – 0.006 |
| Intraspecific | 0.007 ± 0.006 | 0.0 – 0.034 |
| Interspecific | 0.033 ± 0.009 | 0.006 – 0.066 |
In all BLAST-based searches against the BOLD database, the sequences with the highest overall similarity to the query sequence were also recovered in the corresponding monophyletic clade.
MycoBank no. MB 343580
Lichen melanophthalma Ramond in Lamarck & de Candolle, Fl. Franc. Ed. 3.2:377 (1805).
Spain, Teruel, Noguera de Albarracín, carretera A-1521 hacia Orihuela del Tremedal, antes de la Peña del Castillo, pista a “Ruta Laguna”, “Peña Aguada”, 30TXK16815, 1590 m alt., on quartzite in a Quercus pyrenaica and Pinus sylvestris forest, 04 October 2010, M. Vivas & J. Rico, Vivas 94, MAF –Lich 16805 (Epitype: MAF-Lich). We were unable to obtain fresh material from the Pic du Midi de Bigorre in the French Pyrenees (location of original type collection of Rhizoplaca melanophthalma). In order to fix the application of the name, we selected a specimen from Teruel, Spain (MAF-Lich 16805) as the epitype. The epitype shares an identical ITS haplotype with specimens collected in Chile, China, Spain, Switzerland, and the USA, and thus appropriately represents the cosmopolitan distribution of Rhizoplaca melanophthalma s.s. The ITS sequence of the epitype is deposited in GenBank under accession no. JX948232.
A morphological description can be found in
Chemistry – Usnic acid (major); usually with psoromic (major), constipatic (minor), dehydroconstipatic (minor), dehydroprotocetraric (minor), and 2’-O-demethylsubpsoromic (minor or trace) acids; occasionally with subpsoromic (minor) and 2’-O-demethylpsoromic (minor) acids.
Supplementary file 2 (terminal label: ‘6604’) &
GenBank no. JX948232.
Phylogenetic notes: Strongly supported as monophyletic lineage in both concatenated multilocus gene tree (ML bootstrap = 95%: posterior probability = 1.0) and the ITS gene topology (ML bootstrap = 91%, this study); and strong speciation probability inferred from multiple loci (BPP speciation probability = 1.0).
In its narrower circumscription, this taxon isknown from Antarctica, Asia (including Central Asia and China), Europe, North and South America. The species has also been recorded from alpine areas in the tropics. However, additional studies are required to verify the identity of these populations. It typically occurs on exposed calcium-poor rock (e.g. basalt, granite, schist), but sometimes on calcium rich sandstone and limestone. It ranges in distribution from arid lowland woodlands into upper montane coniferous forests and the lower portions of the alpine tundra.
. See supplementary file 1.
MycoBank no. MB 803475
USA, Nevada, White Pine County, on Cave Mountain, north of antenna site, Humboldt-Toiyabe National Forest, 39.1734°N, 114.6130°W, on basalt, 3150 m alt., July 2010, S. D. Leavitt & Larry L. St. Clair BRY-C55076 (holotype BRY).
Consists of specimens recovered within ‘clade IVa’ in
Chemistry – Usnic (major), constipatic (minor), dehydroconstipatic (minor), 2’-O-demethylpsoromic (minor), and 2’-O-demethylsubpsoromic (minor or trace) acids; usually with psoromic acid (major); and occasionally with dehydroprotocetraric acid (minor).
GenBank Nos. HM577307 (ITS), HM577081 (IGS), HM577210 (group I intron), HM577441 (MCM 7), and HM576952 (β-tubulin).
Phylogenetic notes: Strongly supported as monophyletic lineage in both concatenated multilocus gene tree (ML bootstrap = 100%: posterior probability = 1.0) and the ITS gene topology (ML bootstrap = 99%, this study); and strong speciation probability inferred from multiple loci (BPP speciation probability ≥ 0.98). Rhizoplaca occulta belongs to a closely related, and well-supported, monophyletic lineage including Rhizoplaca paralis, Rhizoplaca polymorpha, Rhizoplaca porterii, and the obligatory vagrant species Rhizoplaca haydenii and Rhizoplaca idahoensis.
Growing usually on exposed calcium-poor rock (e.g. basalt, granite, schist) in pinyon-juniper woodlands but also occurs free on soil. So far known only from collections in western North America. Rhizoplaca occulta included a total of five individuals from Idaho (3 individuals), Nevada (1), and Utah (1), USA, and included GenBank accessions identified as Rhizoplaca cerebriformis ined. (AF159942) and Rhizoplaca subidahoensis ined. (AF159944).
The name is derived from the Latin “occultus, ” meaning hidden, and refers to the fact that this species was hidden within the phenotypically circumscribed taxon Rhizoplaca melanophthalma sensu lato.
See supplementary file 1.
MycoBank no. MB 803476
USA, Utah, Sevier County, Thousand Lakes Mountain, north of ‘Flat Top’, 38.5111°N, 111.4732°W, on basalt, 2875 m alt., October 1997, Lyndon D. Porter BRY-C55077 (holotype BRY).
Morphologically similar to Rhizoplaca melanophthalma sensu stricto, but consists of specimens recovered within ‘clade IVb’ in
Chemistry– Usnic (major) and psoromic (major) acids; usually with constipatic (minor), dehydroconstipatic (minor), dehydroprotocetraric (minor), lecanoric (major), orsellinic (minor), and subpsoromic (minor) acids; occasionally with gyrophoric (trace), 2’-O-demethylsubpsoromic (minor or trace) and 2’-O-demethylpsoromic (minor) acids.
GenBank Nos. HM577308 (ITS), HM577082 (IGS), HM577211 (group I intron), HM577442 (MCM 7), and HM576953 (β-tubulin)
Phylogenetic notes: Strong to moderate support as monophyletic lineage in both concatenated multilocus gene tree (ML bootstrap = 83%: posterior probability = 0.93), and with weak statistical support in the ITS gene topology (ML bootstrap = 67%, this study); and strong speciation probability inferred from multiple loci (BPP speciation probability = 1.0). Rhizoplaca parilis belongs to a closely related, and well-supported, monophyletic lineage including Rhizoplaca occultum, Rhizoplaca polymorpha, Rhizoplaca porterii, and the obligatory vagrant species Rhizoplaca haydenii and Rhizoplaca idahoensis.
This species usually occurs on exposed calcium-poor rock (e.g. basalt, granite, schist), but sometimes on calcium rich sandstone and limestone. Its habitat ranges from pinyon-juniper woodlands to montane coniferous forests and the lower portions of alpine tundra. This taxon iscurrently known from Asia (including Central Asia and China), Europe, and North and South America
The specific epithet is chosen from the Latin parilis, meaning equivalent, like, or similar, in reference to the morphological similarity between the new species and the other species within the Rhizoplaca melanophthalma species-complex.
See supplementary file 1.
MycoBank no. MB 803477
USA, Idaho, Owyhee County, 43.3202°N, 116.9795°W, 1291 m alt., 04 July 2008, S. D. Leavitt, H. C. Leavitt & J.H. Leavitt BRY-C55093 (holotype BRY).
Rhizoplaca polymorpha consists of specimens recovered within ‘clade IVc’ in
Chemistry– Usnic (major), constipatic (minor), dehydroconstipatic (minor), 2’-O-demethylsubpsoromic (minor or trace) and 2’-O-demethylpsoromic (minor) acids; occasionally with dehydroprotocetraric (minor) and psoromic acid (major).
GenBank Nos. HM577324 (ITS), HM577097 (IGS), HM577227 (group I intron), HM577458 (MCM 7), and HM576968 (β-tubulin).
Phylogenetic notes: Strongly supported as monophyletic lineage in both concatenated multilocus gene tree (ML bootstrap = 82%: posterior probability = 1.0), and weak statistical support in the ITS gene topology (ML bootstrap = 66%, this study); and strong speciation probability inferred from multiple loci (BPP speciation probability ≥ 0.97). Rhizoplaca polymorpha belongs to a closely related, and well-supported, monophyletic lineage including Rhizoplaca occulta, Rhizoplaca parilis, Rhizoplaca porterii, and the obligatory vagrant species Rhizoplaca haydenii and Rhizoplaca idahoensis.
Currently only known from collections in western North America. Its habitat includes pinyon-juniper woodlands and montane coniferous forests, but unattached forms are also known from the McBride Creek Badlands in Western Idaho.
The specific epithet was selected based on the morphologically polymorphic forms within this species, including both umbilicate and vagrant forms.
See supplementary file 1.
MycoBank no. MB 803478
USA, Utah, Wayne County, Thousand Lakes Mountain, vicinity of ‘Flat Top’, near summit, 38.4432°N, 111.4703°W, on basalt, 3400 m alt., October 1997, Lyndon D. Porter BRY-C55096 (holotype BRY).
Morphologically similar to Rhizoplaca melanophthalma sensu stricto, but consists of specimens recovered within ‘clade IVd’ in
Chemistry– Usnic acid (major); usually with psoromic (major), constipatic (minor), dehydroconstipatic (minor), dehydroprotocetraric (minor), subpsoromic (minor), demethylpsoromic (minor), and 2’-O-demethylsubpsoromic (minor or trace) acids.
GenBank Nos. HM577327 (ITS), HM57710 (IGS), HM577461 (MCM 7), and HM576971 (β-tubulin).
Phylogenetic notes: A monophyletic lineage in both concatenated multilocus gene tree with weak statistical support (ML bootstrap < 50%; posterior probability < 0.5), and with strong statistical support in the ITS gene topology (ML bootstrap = 94%, this study); and strong speciation probability inferred from multiple loci (BPP speciation probability ≥ 0.97). Rhizoplaca porterii belongs to a closely related, and well-supported, monophyletic lineage including Rhizoplaca occulta, Rhizoplaca parilis, Rhizoplaca porterii, and the obligatory vagrant species Rhizoplaca haydenii and Rhizoplaca idahoensis.
This species usually occurs on exposed calcium-poor rock (e.g. basalt, granite, schist), but sometimes on calcium rich sandstone and limestone. Its habitat ranges from pinyon-juniper woodland into montane coniferous forests and lower alpine tundra. This taxon iscurrently known only from the western USA (Idaho and Utah).
The new taxon is named in honor ofDr. Lyndon D. Porter, whose research on Rhizoplaca melanophthalma proved invaluable to the present work.
See supplementary file 1.
MycoBank no. MB 803479
USA, Utah, Wayne County, Thousand Lakes Mountain, vicinity of ‘Flat Top’, near summit, 38.4366°N, 111.4677°W, on basalt, 3270 m alt., October 1997, Lyndon D. Porter BRY-C55061 (holotype BRY).
Morphologically similar to Rhizoplaca melanophthalma sensu stricto, but consists of specimens recovered within ‘clade III’ in
Chemistry– Usnic (major), psoromic (major), subpsoromic (minor), and 2’-O-demethylsubpsoromic (minor or trace) acids; usually with constipatic (minor) and 2’-O-demethylpsoromic (minor) acids; and occasionally with dehydroconstipatic (minor) and dehydroprotocetraric (minor) acids.
GenBank Nos. HM577282 (ITS), HM577058 (IGS), HM577187 (group I intron), HM577416 (MCM 7), and HM576927 (β-tubulin).
Phylogenetic notes: A monophyletic lineage in both concatenated multilocus gene tree with strong statistical support (ML bootstrap = 100%; posterior probability 1.0), and with strong statistical support in the ITS gene topology (ML bootstrap = 100%, this study); and high speciation probability inferred from multiple loci (BPP speciation probability = 1.0).
Found growing only on sun-exposed basalt boulders in subalpine meadows in southwestern USA. Currently known only from subalpine habitats on the Aquarius Plateau in southern Utah, USA.
The new taxon is named in honor ofthe late Dr. Sam Shushan, a pioneer in western North American lichenology.
See supplementary file 1.
In this study we described five new species within Rhizoplaca melanophthalma s.l. Our results indicate that a molecular-based approach for specimen identification in the common lichen-forming Rhizoplaca melanophthalma species-complex can effectively assign individuals from cosmopolitan populations to previously circumscribed ‘candidate’ species (
In spite of the limitations in delimiting taxa using molecular data, the effective use of genetic data appears to be essential to appropriately and practically identify natural groups in some phenotypically cryptic lichen-forming fungal lineages (
Within the Rhizoplaca melanophthalma species-complex, DNA barcoding can be performed in a variety of ecological, bio-monitoring and population genetic studies in order to quickly sort specimens into genetically divergent groups. In Rhizoplaca melanophthalma s.l., this barcode application for specimen identification may provide a valuable framework for assessing biogeographic patterns, bio-monitoring research, and prove to be an important tool in making critical conservation-related decisions. The application of molecular-based identification could also be used as a way for both specialists and non-specialists alike to discriminate species that are otherwise difficult to identify, making specimen identification more accessible and more accurate at the same time. In spite of the progress in recognizing independent species-level lineages within Rhizoplaca melanophthalma s.l., high intraspecific distances within Rhizoplaca melanophthalma sensu stricto, suggest additional species-level lineages may potentially be hidden within this lineage. In order to address this question, we are currently developing novel genetic markers (i.e. microsatellites) specific for this group in order to assess population structure and gene flow within this broadly distributed species.
Using ITS sequence data, specimens within the Rhizoplaca melanophthalma species group can be identified by means of DNA barcoding using the publicly available database in BOLD (http://www.boldsystems.org/). Based on our broad intercontinental sampling, it appears that Rhizoplaca melanophthalma s.l. specimens can be accurately identified to species using the BLAST-based identification tool for fungi in BOLD. This provides an objective approach for a broad array of researchers to accurately identify species within this group using ITS sequence data from their collections, regardless of their level of taxonomic expertise.
Although in some cases the ITS region has been shown to be effective for molecular identification using DNA barcoding (
While in some cases data have supported the taxonomic use of secondary metabolic characters for delimiting lichen taxa (
As molecular sequence data become more readily available, they will allow us to better understand the diversity of lichenized fungi. Their use in identifying species will become increasingly important and routinely applied. Other disciplines such as ecology, conservation and physiology will benefit from a more objectively based species circumscription, enabling us to interpret distribution and ecological patterns better and more accurately monitor environmental disturbance and climate change.
We thank Roger Rosentreter, Johnathon Fankhauser, Leigh Johnson, Christopher Jones, Dean Leavitt, Hailey Leavitt, Jackson Leavitt, Lyndon Porter, Monica Proulx, and Peter Ririe for invaluable assistance with this research. This study was supported, in part, by funds from Brigham Young University graduate mentoring and graduate research fellowship awards, the USDA National Forest Service, the Negaunee Foundation and the National Science Foundation (DEB-0949147). PKD thanks to the Ministerio de Ciencia e Innovación, Spain for financial support (CGL2010-21646/BOS, RYC02007-01576). SPO thanks the JAE-Doc program (CSIC) for financial support.
Supplementary Table S1. (doi: 10.3897/mycokeys.7.4508.app1) Microsoft Excel Document (xls).
Explanation note: Collection information for all Rhizoplaca melanophthalma sensu lato specimens included in the present study. Modified from Leavitt et al. (2013).
Supplementary Fig S1. (doi: 10.3897/mycokeys.7.4508.app2) File format: Adobe Portable Document Format (pdf).
Explanation note: Maximum likelihood ITS gene tree of the 240 sampled Rhizoplaca melanophthalma sensu lato specimens. Bootstrap support indicated at nodes. Modified from Leavitt et al. (2013).