(C) 2012 Imke Schmitt. This is an open access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC-BY), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
For reference, use of the paginated PDF or printed version of this article is recommended.
The lichen-forming genus Pertusaria under its current circumscription is polyphyletic and its phylogenetic affiliations are uncertain. Here we study the species of the genera Pertusaria and Varicellaria which containlecanoric acid as major constituent, have disciform apothecia, strongly amyloid asci, non-amyloid hymenial gel, 1-2-spored asci, and 1- or 2-celled ascospores with thick, 1-layered walls. We infer phylogenetic relationships using maximum likelihood and Bayesian analyses based on four molecular loci (mtSSU, nuLSU rDNA, and the protein-coding, nuclear RPB1 and MCM7 genes). Our results show that the lecanoric acid-containing species form a well-supported, monophyletic group, which is only distantly related to Pertusaria s.str. The phylogenetic position of this clade is unclear, but placement in Pertusaria s.str. is rejected using alternative hypothesis testing. The circumscription of the genus Varicellaria is enlarged to also include species with non-septate ascospores. Seven species are accepted in the genus: Varicellaria culbersonii (Vězda) Schmitt & Lumbsch, comb. nov., Varicellaria hemisphaerica (Flörke) Schmitt & Lumbsch, comb. nov., Varicellaria kasandjeffii (Szatala) Schmitt & Lumbsch, comb. nov., Varicellaria lactea (L.) Schmitt & Lumbsch, comb. nov., Varicellaria philippina (Vain.) Schmitt & Lumbsch, comb. nov., Varicellaria rhodocarpa (Körb.) Th. Fr., and Varicellaria velata (Turner) Schmitt & Lumbsch, comb. nov. A key to the species of Varicellaria is provided.
Agyriales, Ascomycota, lichen-forming fungi, molecular phylogeny, Ostropomycetidae, Pertusaria, Pertusariales, taxonomy
Generic classifications in lichen-forming fungi have changed dramatically since the introduction of molecular data. Numerous genera have been shown to be polyphyletic or nested within larger genera (e.g.,
We assembled a four-locus data set consisting of mtSSU rDNA, nuLSU rDNA, and the protein-coding genes RPB1 and MCM7. The alignment contained 31 species. Specimens and sequences used for molecular analyses are listed in Table 1. Two sequences of Parmeliaceae (Lecanoromycetes) were used as outgroup, since Lecanoromycetes was shown to be a sister-group of Ostropomycetidae to which Pertusariales belongs (
Species and sequences used in this study. New sequences are indicated in bold.
| Name | Phylogenetic lineage | Family | nuLSU | mtSSU | 1RPB | 7MCM |
|---|---|---|---|---|---|---|
| Varicellaria culbersonii* | Varicellaria | ? | JX101871 | JX101873 | JX101875 | JX101874 |
| Varicellaria hemisphaerica | Varicellaria | ? | AF381556 | AF381563 | DQ902341 | GU980998 |
| Varicellaria lactea | Varicellaria | ? | AF381557 | AF381564 | DQ870971 | GU981000 |
| Varicellaria rhodocarpa | Varicellaria | ? | AF381559 | AF381569 | N/A | N/A |
| Varicellaria velata | Varicellaria | ? | AY300855 | GU980981 | DQ870982 | GU981005 |
| “Pertusaria” amara | Variolaria | ? | AF274101 | AY300900 | DQ870965 | GQ272423 |
| “Pertusaria” corallina | Variolaria | ? | AY300850 | AY300901 | DQ870967 | GU980997 |
| “Pertusaria” scaberula | Variolaria | ? | AF274099 | AF431959 | DQ870980 | GU981003 |
| “Pertusaria” subventosa | Variolaria | ? | AY300854 | AY300905 | DQ870981 | GU981004 |
| Circinaria contorta | Megasporaceae | DQ986782 | DQ986876 | DQ986852 | GU980989 | |
| Circinaria hispida | Megasporaceae | DQ780305 | HM060722 | DQ870933 | DQ780273 | |
| Lobothallia radiosa | Megasporaceae | DQ780306 | DQ780274 | DQ870954 | GQ272397 | |
| Ochrolechia parella | Ochrolechiaceae | AF274097 | GU980977 | DQ870959 | GQ272421 | |
| Ochrolechia subpallescens | Ochrolechiaceae | GU980985 | GU980978 | GU981008 | GU980994 | |
| Ochrolechia upsaliensis | Ochrolechiaceae | GU980986 | GU980979 | GU981009 | GU980995 | |
| Coccotrema cucurbitula | Coccotremataceae | AF274092 | AF329161 | DQ870939 | GU980990 | |
| Coccotrema maritimum | Coccotremataceae | AF329164 | AF329163 | N/A | GU980991 | |
| Coccotrema pocillarium | Coccotremataceae | AF274093 | AF329166 | DQ870940 | GU980992 | |
| Gyalectaria diluta | Coccotremataceae | GU980982 | GU980974 | N/A | N/A | |
| Gyalectaria gyalectoides | Coccotremataceae | GU980983 | GU980975 | GU981006 | GU980993 | |
| Gyalectaria jamesii | Coccotremataceae | GU980984 | GU980976 | GU981007 | N/A | |
| Thamnolia vermicularis | Icmadophilaceae | AY961599 | AY853345 | DQ915599 | N/A | |
| Icmadophila ericetorum | Icmadophilaceae | DQ883694 | DQ986897 | DQ883723 | N/A | |
| Dibaeis baeomyces | Icmadophilaceae | AF279385 | AY300883 | DQ842011 | N/A | |
| Agyrium rufum | Agyriaceae | EF581826 | EF581823 | EF581822 | GU980988 | |
| Miltidea ceroplasta** | Miltideaceae | HQ391558 | HQ391557 | JQ900620 | N/A | |
| Pertusaria hermaka*** | Pertusaria s. str. | Pertusariaceae | DQ780334 | DQ780299 | JX101872 | GU980999 |
| Pertusaria paramerae | Pertusaria s. str. | Pertusariaceae | DQ780328 | GU980980 | GU981012 | GU981001 |
| Pertusaria pustulata | Pertusaria s. str. | Pertusariaceae | DQ780332 | DQ780297 | GU981013 | GU981002 |
| Parmeliopsis hyperopta | outgroup | Parmeliaceae | AY607823 | AY611167 | EF092142 | GQ272426 |
| Everniopsis trulla | outgroup | Parmeliaceae | EF108290 | EF108289 | EF105429 | GQ272396 |
*source: Costa Rica, R. Lücking 15424 (F)
**source: Australia, H.T. Lumbsch 20004b, S. Parnmen & T. Widhelm (F)
***source: Australia, A. Mangold, 22 March 2005 (MIN)
Sequence alignments and phylogenetic analysis
We assembled partial sequences using Geneious Pro 5.4.3 (
We analyzed the alignments using maximum likelihood (ML) and Bayesian inference. To test for potential conflict between data sets, we performed ML analyses on the individual alignments and examined the trees for conflicts supported by 75% bootstrap support. ModelTest (
The ML analysis of the concatenated alignment was performed with the program RAxML (
As in previous studies (e.g.
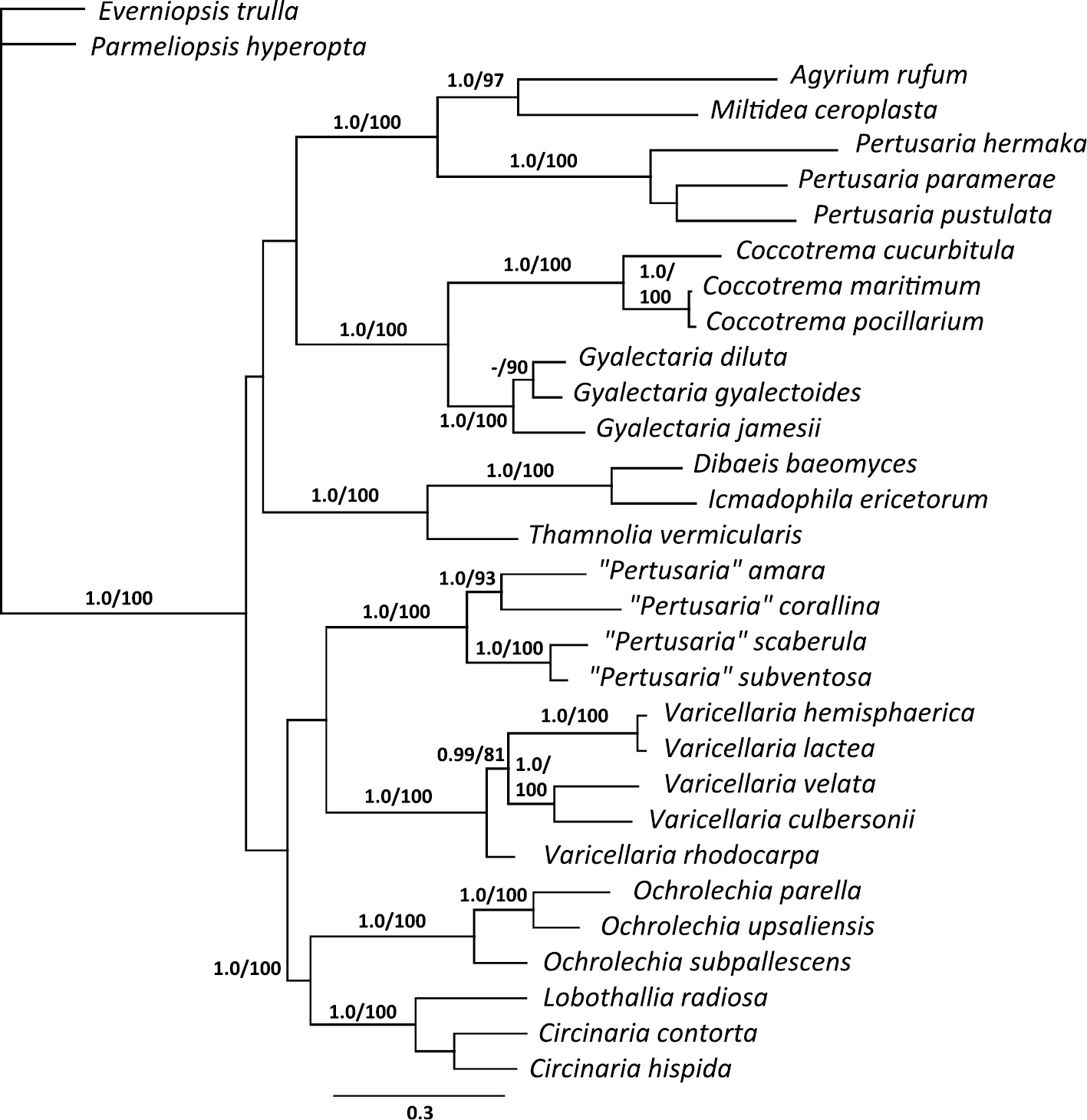
We obtained six new sequences indicated in Table 1. The combined alignment of the nuLSU, mtSSU rDNA, RPB1, and MCM7 included 2790 unambiguously aligned nucleotide position characters, 1226 of which were variable. The single locus ML topologies did not show any conflicts and hence a concatenated analysis was performed. The maximum likelihood tree did not contradict the Bayesian tree topologies and thus only the majority-rule consensus tree of the Bayesian tree sampling is shown here (Fig. 1). In the phylogenetic tree, species of the Varicellaria-group form a strongly supported monophyletic group, including Pertusaria culbersonii. The Varicellaria-group is sister to the Variolaria-group, but this relationship lacks support. The genus Ochrolechia is a well-supported sister-group to Megasporaceae (Circinaria and Lobothallia), and this clade is sister to the Varicellaria- and Variolaria-groups, but again this relationship lacks support. Agyrium and Miltidea form a supported sister-group, which is strongly supported sister to the well-supported, monophyletic Pertusaria s.str. The well-supported, monophyletic genera Coccotrema and Gyalectaria have a well-supported sister-group relationship. The sister-group relationship of Coccotremataceae and the clade including Agyrium, Miltidea, and Pertusaria s.str. lacks support. A placement of the Varicellaria clade in Pertusaria s.str. is rejected significantly (p≤0.001 in both tests) using alternative hypothesis testing.
Phylogeny of pertusarialean fungi based on mtSSU, nuLSU, RPB1 and MCM7 sequences. This is a 50% majority rule consensus tree based on 14, 000 trees from a Bayesian analysis. Values above the branches are posterior probabilities/ML bootstrap support (ML based on 2000 replicates).
The current study confirms previous results on the polyphyly of Pertusaria (
We will address the issue of the phylogeny and classification of the species-rich Variolaria-group in the future using an extended and geographically balanced taxon sampling. Our study shows that additional, molecular markers will be necessary to elucidate the phylogenetic relationships of major clades within Pertusariales (incl. Agyriales) (
Varicellaria microsticta Nyl. Mém. Soc. Imp. Sci. Nat. Cherbourg 5: 119. 1858. [=Varicellaria rhodocarpa (Körb.) Th.Fr.]
=Clausaria Nyl. Annls Sci. Nat., Bot., sér. 4 15: 45. 1861.
Type species. Clausaria fallens Nyl., Ann. Sci. Nat., Bot., sér. 4 15: 45. 1861. [=Varicellaria velata (Turner) Schmitt & Lumbsch]
The genus in its enlarged circumscription includes species with disciform ascomata, non-amyloid hymenial gel, strongly amyloid, 1-2-spored asci, and 1- or 2-celled ascospores with thick, 1-layered walls. All species contain lecanoric acid, and may also contain lichexanthone or variolaric acid. Currently, we accept seven species in this genus. The accepted names and authorities are listed below.
Pertusaria culbersonii Vězda. Lich. sel. exs. 60: 4 (no. 1487). 1977. Type. Costa Rica, San José, Cerro de la Muerte, 3330m alt., 1976, on soil, W.L. Culberson 13195J (holotype PRA-V).
Variolaria hemisphaerica Flörke. Deutsche Lich. 2: 6. 1815. Type. Germany, Berlin [Flörke, Deutsche Lichenen exs. 29] (isotype BM).
Pertusaria hemisphaerica (Flörke) Erichsen. Hedwigia 72: 85. 1932.
Pertusaria kasandjeffii Szatala. Magy. Bot. Lapok 29: 83. 1930. Type. Bulgaria, Cepelarska planina, in monte Turluka, par Pamsakli, 1500m alt., 6.1929, Szatala (isotype HBG-1233).
This species is only known from a few localities in Bulgaria and Romania (
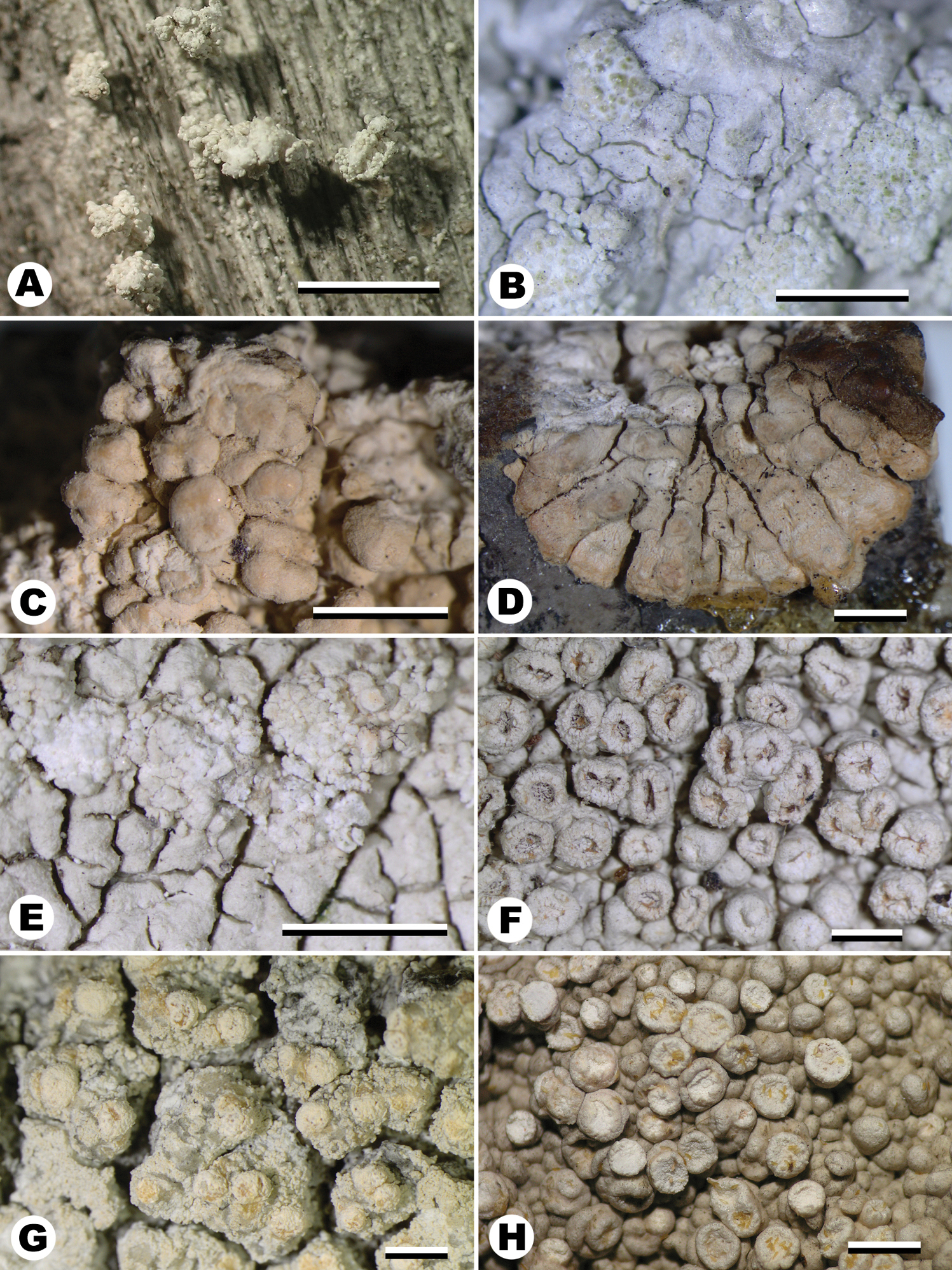
The species of Varicellaria. A Varicellaria culbersonii. Costa Rica, Buck 44182 (F) B Varicellaria hemisphaerica. Germany, 15.4.2004, Schmitt (FR) C, D Varicellaria kasandjeffii. Isotype. Bulgaria, Cepalarska planina: in monte Turluka, par Pasmakali, 1500 m, 9.6.1929, Szatala (HBG-1233) E Varicellaria lactea. Spain, Schmitt 5.6.2003 (FR) F Varicellaria philippina. Holotype. Philippines, Mindanao Dist. Lanao, Camp Keithley by lake Lanao, Sept. 1907, M.S. Clemens, (TUR-V-0006709) G Varicellaria rhodocarpa. Sweden, Printzen 6908 (FR) H Varicellaria velata. Colombia, Moncada & Davila 1537 (F). Scale bar: 1mm. Images were taken with an Olympus SC30 camera under an Olympus SZX7 stereomicroscope.
Lichen lacteus L., Mant. Pl. 1: 132. 1767. Type. Sweden, Västergötland, Mularp, 6.08.1922, Vrang [=Malme, Lich. Suec. Exs. 848] (neotype UPS, designated by
Lepra lactea (L.) F.H.Wigg. Prim. fl. Holsat.: 97. 1780. Variolaria lactea (L.) Pers. Ann. Bot. 1: 24. 1794. Psora lactea (L.) P.Gaertn., G.Mey. & Scherb. Ökonom.-techn. Fl. Wetterau 3: 214. 1801. Zeora lactea (L.) Arnold. Flora, Jena 53: 214. 1870. Pertusaria lactea (L.) Arnold. Verh. zool.-bot. Ges. Wien 22: 283. 1872. Ochrolechia lactea (L.) Matzer & Hafellner. Bibl. Lichenol. 37: 101. 1990.
Mycobank: MB 800589
Pertusaria philippina Vain. Philipp. J. Sci., C, Bot. 8: 131. 1913. Type. Philippines, Mindanao, Lanao, Castra Keithley at Lake Lanao, 1907, Clemens 1302 (holotype TUR-V 6391!).
This species is only known from the Philippines (
Pertusaria rhodocarpa Körb. Syst. lich. germ.: 384. 1855.
Varicellaria microsticta Nyl. Mém. Soc. Imp. Sci. Nat. Cherbourg 5: 119. 1858. Varicellaria kemensis Räsänen. Ann. Soc. zool.-bot. Fenn. Vanamo 3: 295. 1926.
Parmelia velata Turner. Trans. Linn. Soc. London 9: 143. 1808. Type. Great Britain, England, Sussex, 1805, Borrer (holotype BM-4109).
Lichen velatus (Turner) Sm. & Sowerby. Engl. Bot. 29: tab. 2062. 1809. Variolaria velata (Turner) Ach. Lich. univ.: 696. 1810. Pertusaria velata (Turner) Nyl. Lich. Scand. (Uppsala): 179. 1861.
Pertusaria conglobata (Ach.) Th.Fr. Lichenogr. Scand. 1: 321. 1871. Variolaria conglobata Ach. Syn. Lich.: 132. 1814.
Pertusaria haematommoides Zahlbr., Feddes Rep. 33: 50. 1933. Type. Taiwan, Rengechi, Asahina 263 (W – holotype!).
Pertusaria obvelata Nyl. Bih. K. svenska Vetensk. Akad. Handl. 3: 1–156. 1888.
| 1a | Ascospores 2-celled, thallus esorediate or rarely sorediate, containing lecanoric acid, growing on soil, detritus or mosses in arctic-alpine habitats of the northern Hemisphere | Varicellaria rhodocarpa |
| 1b | Ascospores 1-celled, thallus esorediate or sorediate, chemistry and habitat various | 2 |
| 2a | Thallus esorediate | 3 |
| 2b | Thallus sorediate | 6 |
| 3a | Thallus thin, coarsely wrinkled to rimose-cracked, containing lecanoric acid, ± lichexanthone, and ± variolaric acid | 4 |
| 3b | Thallus thick, bullate, apothecia rare or unknown, when present 1-1.5 mm in diam., lacking lichexanthones, Neotropical or restricted to eastern Europe | 5 |
| 4a | Asci 1-spored, cosmopolitan | Varicellaria velata |
| 4b | Asci 2-spored, so far only known from Philippines and Papua New Guinea | Varicellaria philippina |
| 5a | Growing on siliceous rocks, known only from the Balkan region of Europe | Varicellaria kasandjeffii |
| 5b | Growing on soil, detritus or mosses, known from high altitudes in Central America | Varicellaria culbersonii |
| 6a | Thallus containing lecanoric acid, on bark, rarely on rocks | Varicellaria hemisphaerica |
| 6b | Thallus containing lecanoric acid and variolaric acid, on rocks, rarely on bark | Varicellaria lactea |
We thank Armin Mangold (Berlin) and Todd Widhelm (Omaha) for collecting material used in this study, and Miklós Bálint (Frankfurt) for assisting with running phylogenetic analyses on the “FUCHS” computing cluster of the Center for Scientific Computing, Frankfurt am Main. Matthias Schultz (Hamburg), Seppo Huhtinen (Turku), and Anton Igersheim (Vienna) kindly sent type material on loan, and Klaus Kalb (Neumarkt) provided literature. This study was funded by ‘LOEWE, Landes-Offensive zur Entwicklung Wissenschaftlich-ökonomischer Exzellenz’ of Hesse’s Ministry of Higher Education, Research, and the Arts. A.S-D. was supported by a stipend from Deutsche Bundesstiftung Umwelt (DBU).