|
||
|
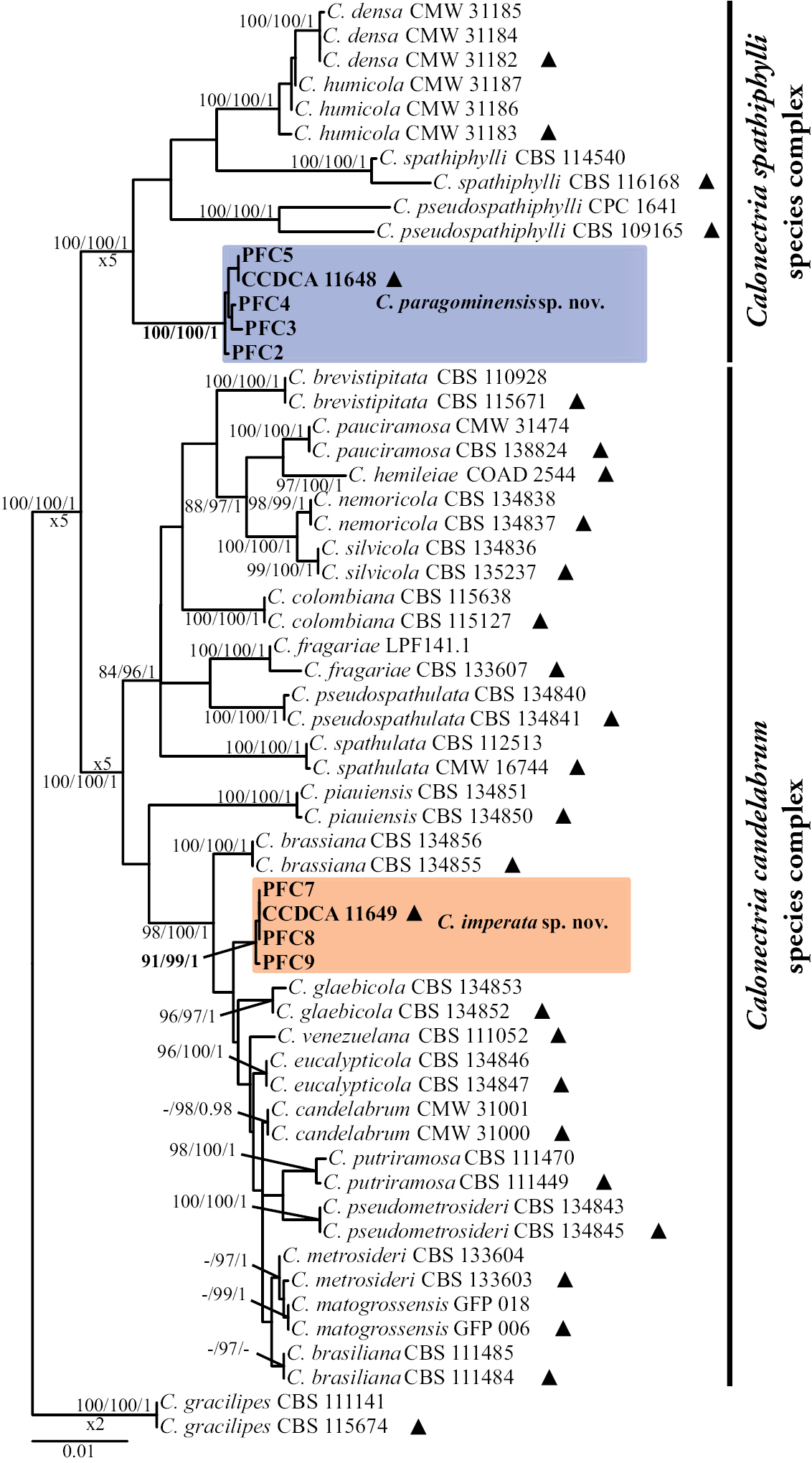
Phylogenetic tree based on maximum likelihood analysis of concatenated act, cmdA, his3, rpb2, tef1 and tub2 gene regions. Bootstrap support values ≥ 80% for maximum parsimony (MP), Ultrafast bootstrap support values ≥ 95% for maximum likelihood (ML), and posterior probability (PP) values ≥ 0.95 from BI analyses are presented at the nodes (MP/ML/PP). Bootstrap values below 80% (MP), 95% (ML) and posterior probabilities below 0.80 are marked with “-”. Ex-type isolates are indicated by “▲”, isolates highlighted in bold were sequenced in this study, and novel species are in blue and orange. C. gracilipes was used as outgroup. The scale bar indicates the number of nucleotide substitutions per site. |